

MAFFT version 7
Better to avoid the Desktop and Documents folders, because these folders may be protected by "Controlled folder access" in "Ransomware protection" in Windows Defender.
When using other tools for extracting, there are rare cases where LF in text files in this package is converted to CRLF. Make sure to keep LF.
| 1 | Copy an input file (fasta format; example) into the mafft-win folder, which was extracted from the package. |
| 2 |
Double-click on |
| 2′ |
Alternatively, to run the signed version, right-click (not double-click) on |
| 3 | Follow the prompts. Filename extension (eg., .txt) must be typed, if any. |
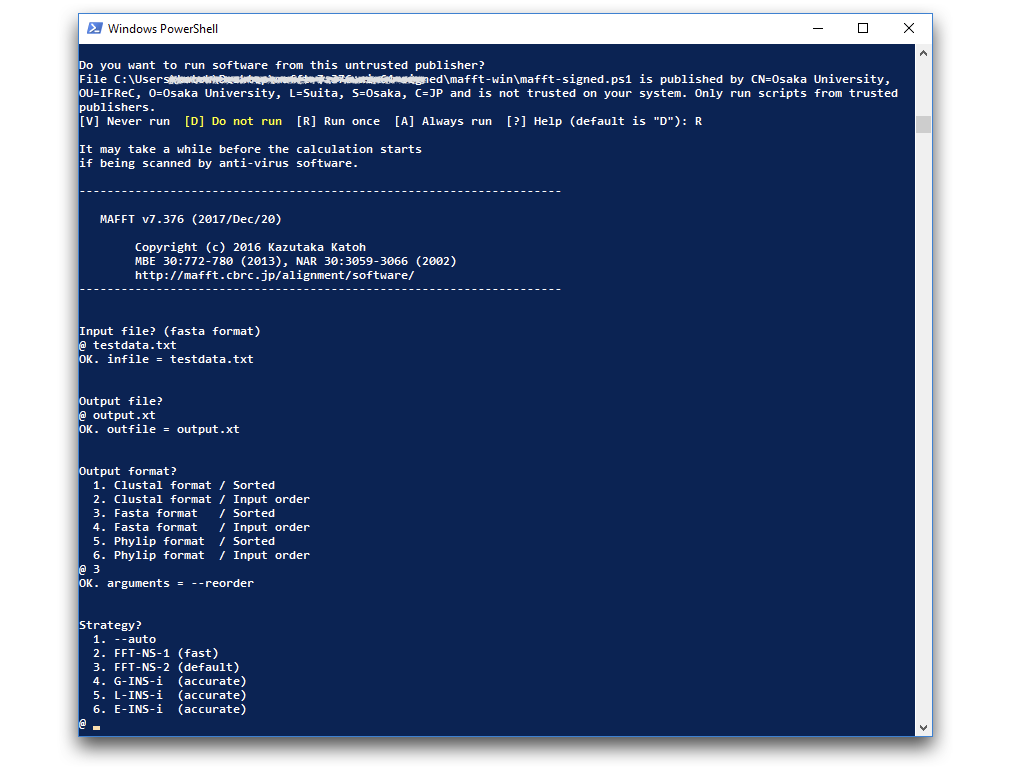
C:\somewhere\mafft-win>mafft.bat --out output.txt --localpair input.txt
This function had a problem in version 7.407 (released 2018/Jul).
Fixed in version 7.409 (2018/Aug).
To run mafft-signed.ps1, the signed version, use Windows PowerShell.
PS C:\somewhere\mafft-win> Set-ExecutionPolicy -Scope CurrentUser AllSigned PS C:\somewhere\mafft-win> .\mafft-signed.ps1 --out output.txt --localpair input.txt PS C:\somewhere\mafft-win> Set-ExecutionPolicy -Scope CurrentUser RestrictedExecutionPolicy is configured before and after running mafft in this case.
call C:\somewhere\mafft-win\mafft.bat --localpair --out output.txt input.txt
cmd.exe /C C:\somewhere\mafft-win\mafft.bat --localpair --out output.txt input.txt
To call the signed script, mafft-signed.ps1,
powershell.exe -ExecutionPolicy AllSigned C:\somewhere\mafft-win\mafft-signed.ps1 --localpair --out output.txt input.txt
There are more efficient ways to directly call binary executables (.exe), instead of a script, in Windows environment. Contact us if necessary.